Contact Us
For additional information about the Computational Lung Physiology Laboratory, Contact the CLPL.
Report a Problem
To report a problem with this website, contact BME Communications or report an accessibility issue.
Since oxidative stress plays a critical role in lung pathophysiology, the Computational Lung Physiology Laboratory (CLPL) is developing molecular fluorescence imaging for probing cellular function in lung cells and tissue. We commonly use an experimental protocol in which the
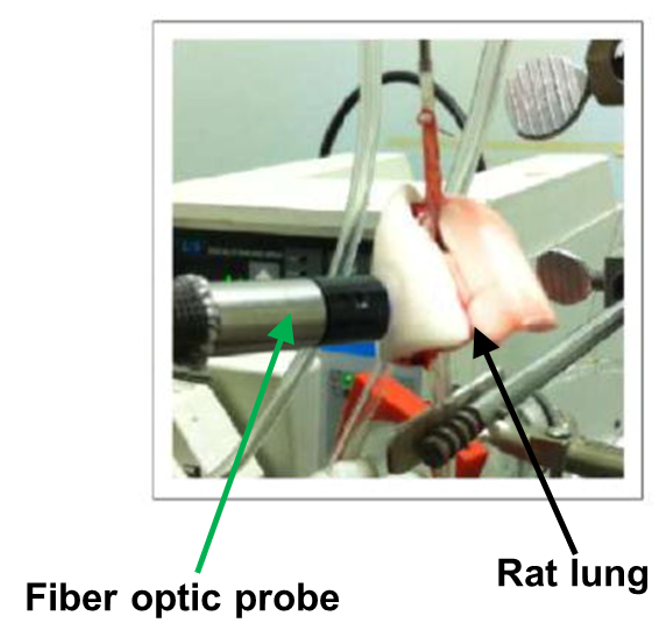 lung of a rodent is removed from the body and connected to a ventilation and perfusion system. We then have the ability to spectrophotometrically measure the concentration of a fluorescent probe entering, exiting, or residing in the lung under different experimental conditions. These data—together with a physiologically-based pharmacokinetic model—are then used to determine the value of key constants associated with particular cellular processes that change with injury. For example, we have developed a robust protocol to measure the rate of hydrogen peroxide (H2O2) production in lungs as an index of oxidative stress and to determine the cellular sources of the measured H2O2using the extracellular probe Amplex Red. We have shown that complex II of the electron transport chain is a potentially important source and/or regulator of mitochondrial H2O2. We are also developing experimental and computational approaches for estimating mitochondrial membrane potential in lungs using the fluorescent probe rhodamine 6G. This approach has potential to provide a quantitative assessment of the effect of injurious conditions on lung mitochondrial function and to evaluate the impact of therapies that target mitochondria.
lung of a rodent is removed from the body and connected to a ventilation and perfusion system. We then have the ability to spectrophotometrically measure the concentration of a fluorescent probe entering, exiting, or residing in the lung under different experimental conditions. These data—together with a physiologically-based pharmacokinetic model—are then used to determine the value of key constants associated with particular cellular processes that change with injury. For example, we have developed a robust protocol to measure the rate of hydrogen peroxide (H2O2) production in lungs as an index of oxidative stress and to determine the cellular sources of the measured H2O2using the extracellular probe Amplex Red. We have shown that complex II of the electron transport chain is a potentially important source and/or regulator of mitochondrial H2O2. We are also developing experimental and computational approaches for estimating mitochondrial membrane potential in lungs using the fluorescent probe rhodamine 6G. This approach has potential to provide a quantitative assessment of the effect of injurious conditions on lung mitochondrial function and to evaluate the impact of therapies that target mitochondria.
Detection of hydrogen peroxide production in the isolated rat lung using Amplex red. (Audi SH, Friedly N, Dash RK, Beyer AM, Clough AV, Jacobs ER) Free Radic Res 2018 Sep;52(9):1052-1062 (PMID: 30175632)
Quantitative optical measurement of mitochondrial superoxide dynamics in pulmonary artery endothelial cells. (Ghanian Z, Konduri GG, Audi SH, Camara AKS, Ranji M) J Innov Opt Health
The Computational Lung Physiology Laboratory has a strong commitment to training the next generation of scientists interested in using state-of-the-art computational techniques to improve human health. Research opportunities are available for undergraduate and graduate (MS and PhD) students alike. For more information on joining our team, Contact the CLPL !