Contact Us
For additional information about the Computational Lung Physiology Laboratory, Contact the CLPL.
Report a Problem
To report a problem with this website, contact BME Communications or report an accessibility issue.
The research methodology at the Computational Lung Physiology Laboratory (CLPL) includes the use of optical and molecular imaging, in combination with computational modeling, for mechanistic interpretation of experimental data and the use of genetically modified rodents and other animal models of human lung disease.
Learn more about projects currently underway in the Computational Lung Physiology Laboratory at the Marquette University and Medical College of Wisconsin Department of Biomedical Engineering.
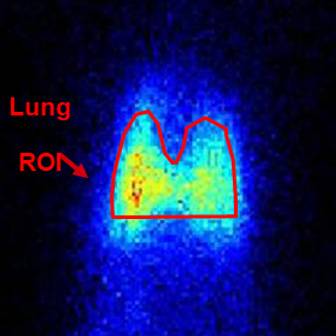 Imaging Lung Injury & Disease
Imaging Lung Injury & DiseaseThe principal investigators at the CLPL have pioneered the use of imaging novel biomarkers as a means of early detection and monitoring of mechanisms and pathways involved in lung injury and disease.
Learn more about Imaging Lung Injury & Disease
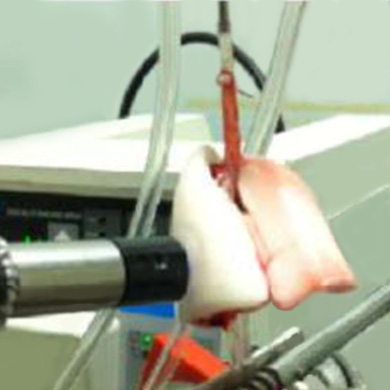 Imaging Cellular Function in Lung Tissue
Imaging Cellular Function in Lung TissueThe CLPL is using molecular fluorescence imaging to probe cellular function in lung cells and tissue. This approach has the potential to provide a quantitative assessment of the effect of injurious conditions on lung mitochondrial function and to evaluate the impact of therapies that target mitochondria.
Learn more about Imaging Cellular Function in Lung Tissue
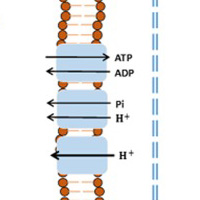 Computational Modeling of Lung Bioenergetics
Computational Modeling of Lung BioenergeticsTo assist with interpreting the complex processes of lung function, the CLPL is using computational modeling as a framework for integrating bioenergetic data at different levels of biological organization.
Learn more about Computational Modeling of Lung Bioenergetics
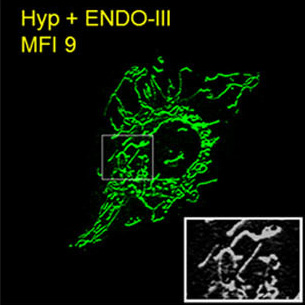 Mitochondrial Dysfunction in Lung Injury
Mitochondrial Dysfunction in Lung InjuryThe CLPL is investigating the molecular basis of hyperoxia-associated mitochondrial-fragmentation in pulmonary endothelial cells and subsequent pulmonary microvascular permeability, complications commonly associated with mechanical ventilation.
Learn more about Mitochondrial Dysfunction in Lung Injury
Research at the Computational Lung Physiology Laboratory is supported on a continual basis by the Marquette University and Medical College of Wisconsin Joint Department of Biomedical Engineering and on a per-project basis by various institutions such as the National Institutes of Health.
The Computational Lung Physiology Laboratory has a strong commitment to training the next generation of scientists interested in using state-of-the-art computational techniques to improve human health. Research opportunities are available for undergraduate and graduate (MS and PhD) students alike. For more information on joining our team, Contact the CLPL !